Learning from deep learning
Can artificial intelligence (AI) read more from a tissue section than even experienced pathologists see under the microscope? In the project Learning from Deep Learning, we investigate how deep neural networks, trained on routine microscopic images of cancer tissue, can predict patient outcomes several years into the future – and what do the models actually "see" when making such assessments. The project is supported by the Research Council of Norway.
What can we learn from better predictions?
The DoMore AI model has demonstrated that deep neural networks can provide highly accurate predictions for conditions such as colorectal cancer (CRC) and prostate cancer (PCa), based solely on H&E-stained sections. In the Learning from Deep Learning project, we investigate
- which tissue structures and cell types drive the model's assessments
- which characteristics are associated with metastases and long-term survival
- how this information can be used in a more transparent manner in the clinic
We achieve this by combining biomarker mapping, explainable artificial intelligence, and multimodal integration.
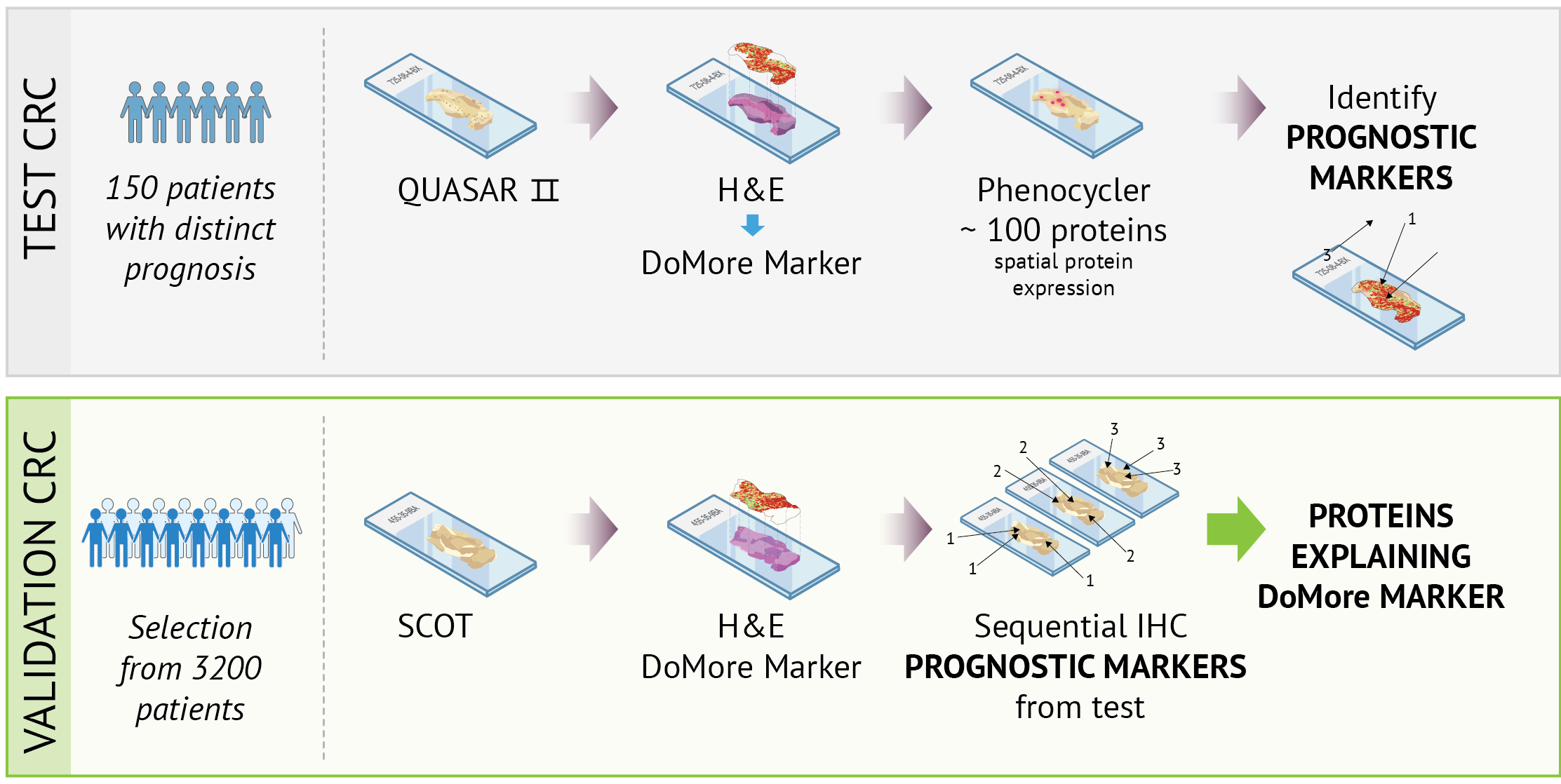
Routine H&E-stained tissue sections, which highlight cells and tissue structures, are used to train the AI model. In the project, these sections are de-stained and re-stained using methods that reveal other types of protein expression. This allows us to link the patterns the model identifies in the H&E image to biomarkers—measurable biological characteristics, such as specific proteins or tissue patterns—that provide insights into diagnosis, disease progression, or treatment efficacy.
We employ spatial proteomics (PhenoCycler-Fusion) and sequential immunostaining to map protein expression in tumour tissue and correlate it with areas that the DoMore model assesses as high- or low-risk. Using multiplex-based analyses, detailed proteomic profiles can be studied directly within the spatial context of the tissue. These data are integrated with morphological analyses and genomic data, enabling us to investigate how tissue architecture, protein expression, and genetic changes collectively contribute to AI-based prognostic markers.
Explainable AI and multimodal integration
We utilise methods within explainable artificial intelligence to identify which tissue characteristics the AI models emphasise. Through analyses of heat maps, internal representations in neural networks, and attention mechanisms, we highlight tissue patterns and microenvironments associated with varying prognoses.
These findings are integrated with biomarker data and genomic analyses in multimodal models. Through cell segmentation and clustering analyses, we study protein expression, cell type composition, and spatial organisation at the single-cell level, examining how molecular and genetic changes are reflected in the tissue architecture that the AI models use in their prognostic assessments.
Towards more comprehensible AI in cancer prognostics
The aim of the Learning from Deep Learning project is to make our AI-based prognostic model biologically understandable and clinically useful. By linking the model's digital markers to concrete biological mechanisms and tissue patterns, the project will provide new insights into what AI actually measures in tumour tissue, thereby laying the groundwork for more precise risk stratification and future AI-based decision support in cancer treatment.